Population Guide¶
The “Population” object in evol is the base container for all your candidate solutions. Each candidate solution (sometimes referred to as a chromosome in the literature) lives inside of the population as an “Individual” and has a fitness score attached as a property.
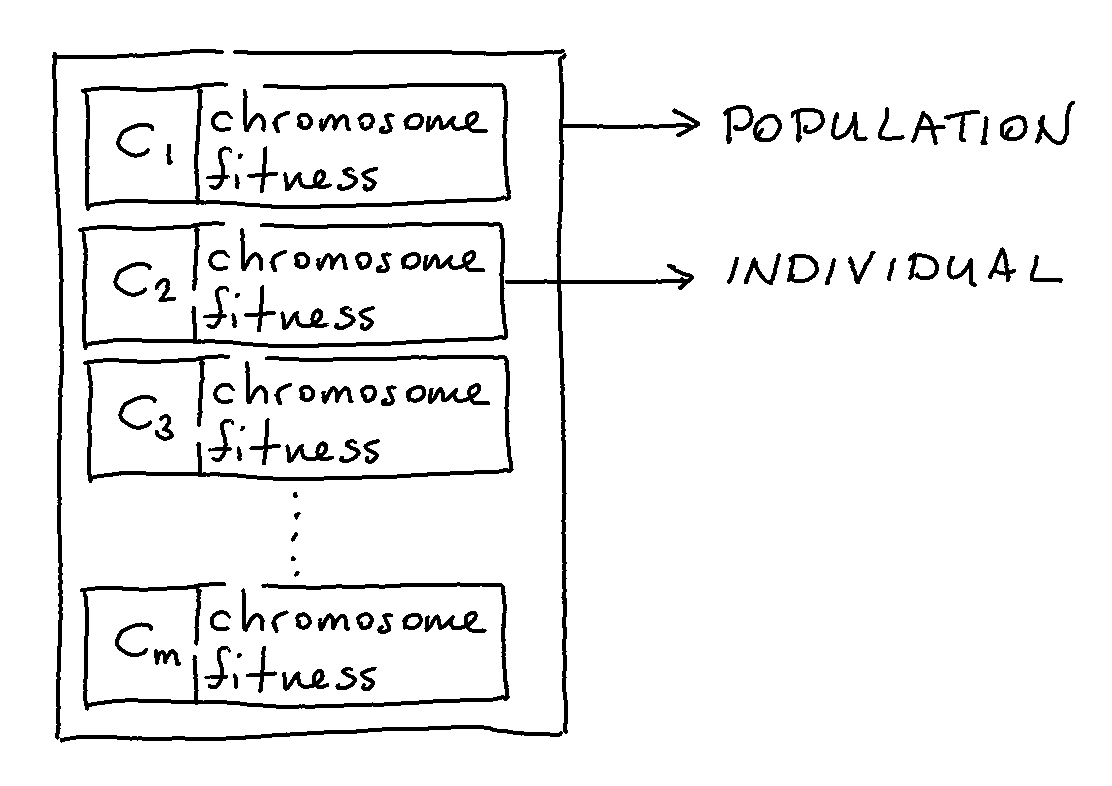
You do not typically deal with “Individual” objects directly but it is useful to know that they are data containers that have a chromosome property as well as a fitness property.
Creation¶
In order to create a population you need an evaluation function and either:
- a collection of candidate solutions
- a function that can generate candidate solutions
Both methods of initialising a population are demonstrated below.
import random
from evol import Population
def create_candidate():
return random.random() - 0.5
def func_to_optimise(x):
return x*2
def pick_random_parents(pop):
return random.choice(pop)
random.seed(42)
pop1 = Population(chromosomes=[create_candidate() for _ in range(5)],
eval_function=func_to_optimise, maximize=True)
pop2 = Population.generate(init_function=create_candidate,
eval_function=func_to_optimise,
size=5,
maximize=False)
Lazy Evaluation¶
If we were to now query the contents of the population object you can use a for loop to view some of the contents.
> [i for i in pop1]
[<individual id:05c4f0 fitness:None>,
<individual id:cdd150 fitness:None>,
<individual id:110e12 fitness:None>,
<individual id:a77886 fitness:None>,
<individual id:8a71e9 fitness:None>]
> [i.chromosome for i in pop1]
[ 0.13942679845788375,
-0.47498924477733306,
-0.22497068163088074,
-0.27678926185117725,
0.2364712141640124]
You might be slightly suprised by the following result though.
> [i.fitness for i in pop1]
[None, None, None, None, None]
The fitness property seems to not exist. But if we call the “evaluate” method first then suddenly it does seem to make an appearance.
> [i.fitness for i in pop1.evaluate()]
[ 0.2788535969157675,
-0.9499784895546661,
-0.4499413632617615,
-0.5535785237023545,
0.4729424283280248]
There is some logic behind this. Typically the evaluation function can be very expensive to calculate so you might want to consider running it as late as possible and as few times as possible. The only command that needs a fitness is the “survive” method. All other methods can apply transformations to the chromosome without needing to evaluate the fitness.
More Lazyness¶
To demonstrate the effect of this lazyness, let’s see the effect of the fitness of the individuals.
First, note that after a survive method everything is evaluated.
> [i.fitness for i in pop1.survive(n=3)]
[0.4729424283280248, 0.2788535969157675, -0.4499413632617615]
If we were to add a mutate step afterwards we will see that the lazyness kicks in again. Only if we add an evaluate step will we see fitness values again.
def add_noise(x):
return 0.1 * (random.random() - 0.5) + x
> [i.fitness for i in pop1.survive(n=3).mutate(add_noise)]
[None, None, None]
> [i.fitness for i in pop1.survive(n=3).mutate(add_noise).evaluate()]
[0.3564375534260752, 0.30990154209234466, -0.5356458732043454]
If you want to work with fitness values explicitly it is good to know about this, otherwise the library will try to be as conservative as possible when it comes to evaluating the fitness function.